Biomaterials Translational ›› 2024, Vol. 5 ›› Issue (1): 21-32.doi: 10.12336/biomatertransl.2024.01.003
• REVIEWS • Previous Articles Next Articles
Zhangjie Li1, Dingyuan Yu1, Chenyang Zhou1, Feifan Wang1, Kangyi Lu1, Yijun Liu1, Jiaqi Xu1, Lian Xuan2, Xiaolin Wang1,2,3,4,*(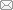 )
)
Received:2024-01-16
Revised:2024-02-29
Accepted:2024-03-14
Online:2024-03-28
Published:2024-03-28
Contact:
Xiaolin Wang, About author:Xiaolin Wang, xlwang83@sjtu.edu.cn.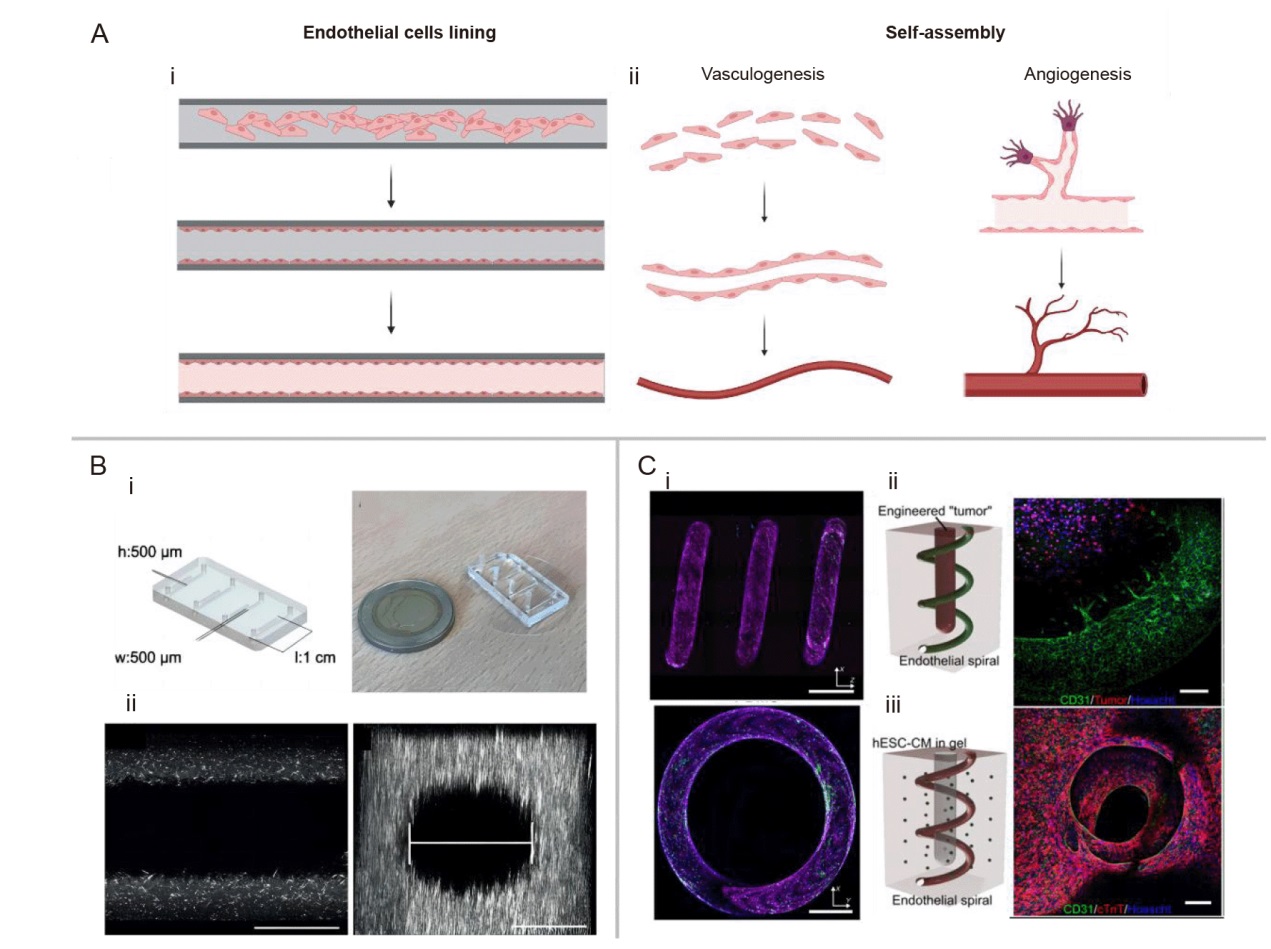
Figure 1. Schematic representation of the principles underlying two methods and top-down EC lining method for constructing vascularised models in vitro. (A) Schematic illustrating two methods for constructing vascularised models in vitro: (i) Principle of the endothelial cell lining method. (ii) Principle of the self-assembly method. Created with BioRender.com. (B) Microvascular system combined with viscous finger patterning technique: (i) Schematic of the microfluidic chip showing dimensions and layout of the microfluidic platform, four straight channels on a single chip with designed parameters 500 μm × 500 μm × 1 cm (w × h × l). (ii) 20× magnification confocal slice of a patterned lumen with 5 mg/mL collagen I. Scale bar: 200 μm. (iii) XZ reconstruction showing the flow field of the scaffold. Scale bar: 200 μm. Reprinted from de Graaf et al.38 (C) Spiral microvascular system: (i) Cross-sectional fluorescence image of the spiral blood vessel. (ii) Spiral vascularised tumour model demonstrating vascular growth towards tumour cells. (iii) Spiral vascularised heart model. Scale bars: 200 μm. Reprinted from Mandrycky et al.24 CM: cardiomyocyte; cTnT: cardiac troponin T; EC: endothelial cell; hESC: human embryonic stem cell.
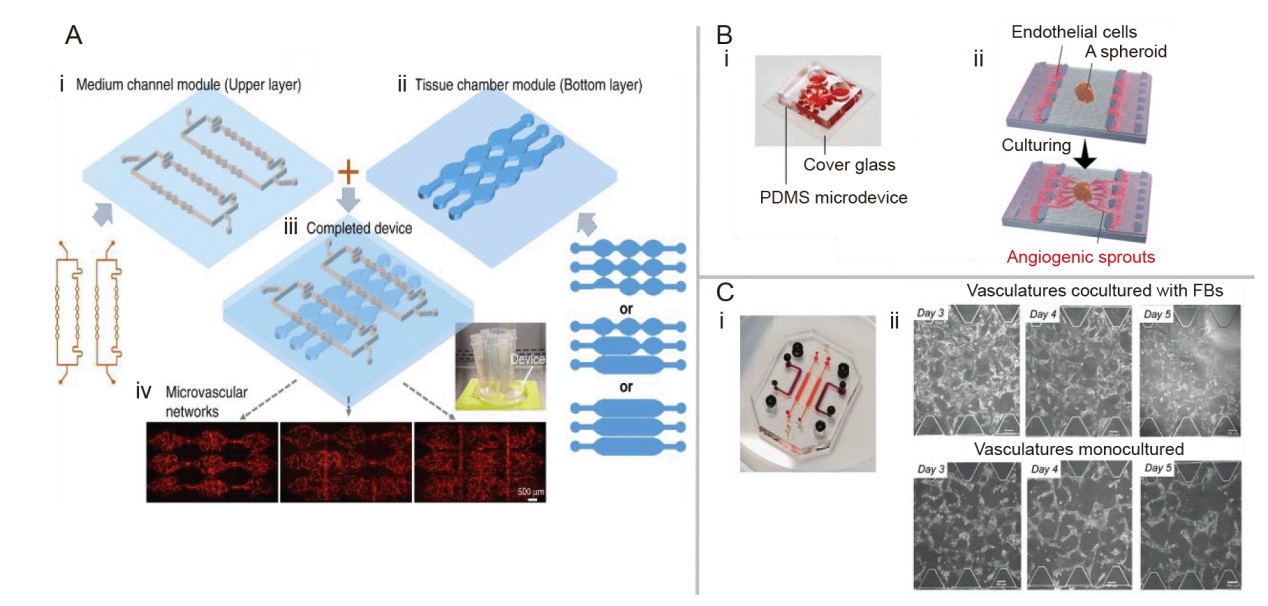
Figure 2. Bottom-up self-assembly vascularisation method for constructing vascularised models in vitro. (A) The modular microfluidic system combines two PDMS layers: (i) Medium channel module (upper layer). (ii) Tissue chamber module (bottom layer). (iii) A completed two-layered device shows the medium channels and tissue chambers in different layers. (iv) Large-scale perfused microvascular networks are generated using different configurations. Reprinted from Yue et al.50 (B) Design diagram and overview of the microfluidic system with perfusion vascular network: (i) Photograph of a microfluidic device. (ii) Schematic diagram showing an overview of assays. Reprinted from Nashimoto et al. 51 Copyright 2017 The Royal Society of Chemistry. Reproduced with permission. (C) Design diagram and representative images of the microfluidic vascular model as a non-contact cell culture device: (i) Photograph of the microfluidic chip used in the study to generate a perfusable vascular network. (ii) Representative images show the comparison of cocultured ECs to monocultured ECs over 3 days. Scale bars: 100 μm. Reprinted from Tan et al.53 EC: endothelial cell; FB: fibroblast: PDMS: polydimethylsiloxane.
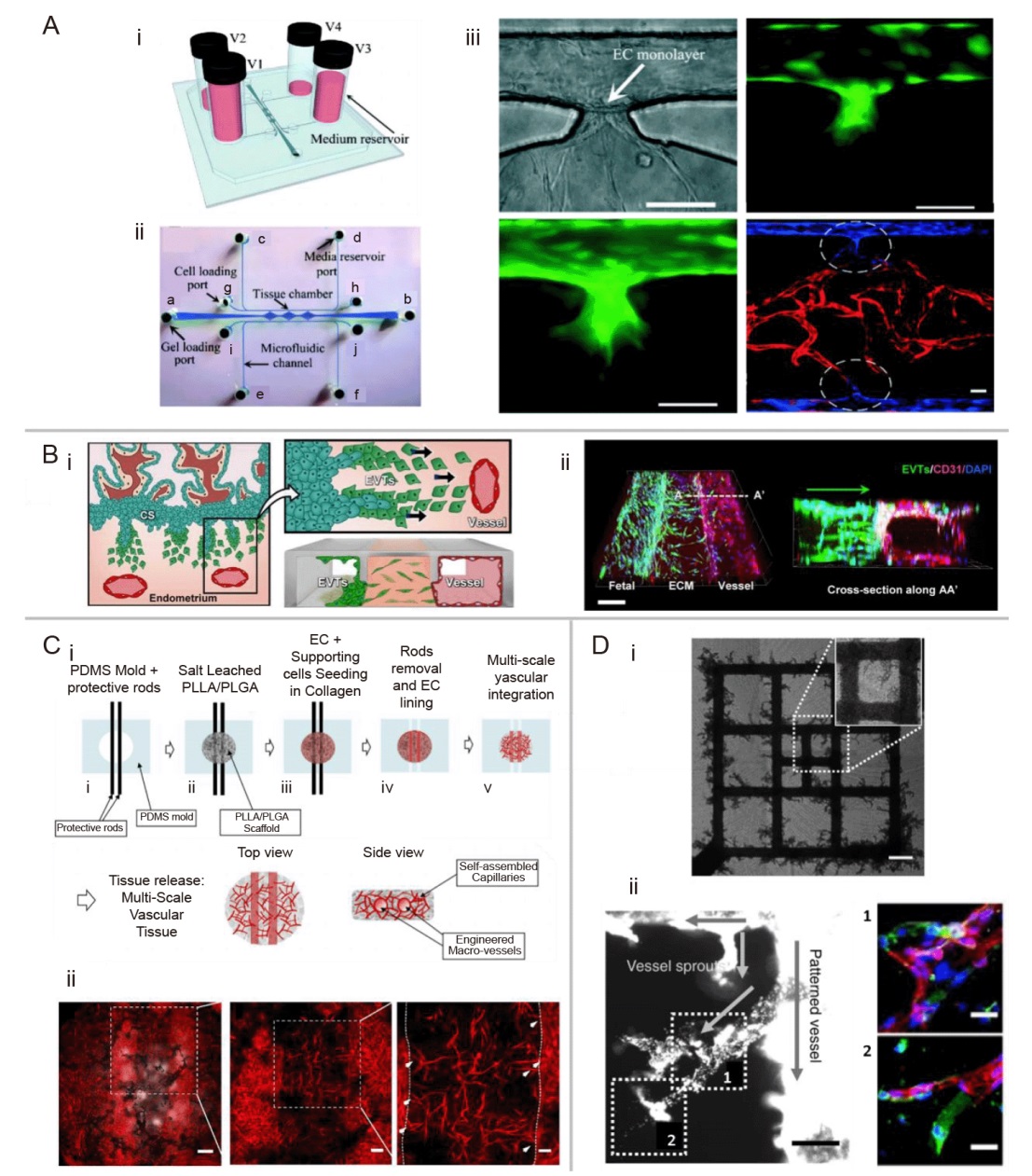
Figure 3. Hybrid method for constructing vascularised models in vitro. (A) Vascularised model construction combining vasculogenesis mechanism and EC lining method: (i) Schematic diagram of the microfluidic chip. The physiological level of mechanical stimulion vasculogenesis was established by filling the reservoirs to different culture medium heights (V1: 23 mm H2O, V2: 8 mm H2O, V3: 18 mm H2O, V4: 3 mm H2O (1 mm H2O = 9.80665 Pa)). (ii) Microchannel design. The gel loading port (a, b), the media reservoir ports (c-f), cell loading port (g-j). (iii) Observation of angiogenesis phenomenon in the microfluidic chip. Scale bars: 100 μm. GFP (green)-labeled indicates the lined ECs at the cell-matrix interface invaded into the 3D fibrin gel and formed microvascular sprouts as early as 24 hours post-lining. From inside the tissue chamber, mCherry-expressing (red) ECs (ECs-mCherry) migrated outward to the laminin-coated microfluidic channel to connect with the lined BFP-labeled (blue) ECs (ECs-BFP) in the channels. The green image below represents more time of microvascular sprouts invasion than the green image above. Reprinted from Wang et al.54 Copyright 2016 The Royal Society of Chemistry. (B) Diagram of the impact of chip implantation on maternal vascular remodelling: (i) Interactions between invading EVTs and maternal blood vessels were modeled in the implantation-on-a-chip. (ii) 3D projection images of EVTs (green) migrating across the ECM hydrogel towards maternal ECs. Scale bar: 200 μm. Reprinted from Park et al.55 (C) MSVT construction: (i) Schematic of the MSVT fabrication process. (ii) Maximum intensity projection of the MSVT, with high magnifications. The middle image is an enlargement of the dashed box in the left image. The right image is an enlargement of the dashed box in the middle image. The white arrows of the right image indicate micro-vessels sprouts and the dashed lines of the right image indicate the macro-vessels borders. Scale bars: 500 μm (left), 100 μm (middle), 50 μm (right). Reprinted from Debbi et al.56 (D) Fluorescent bead perfusion of engineered microvessels (μVs) and citrated whole-blood perfusion in μVs: (i) Brightfield stitched large image of red blood cell-filled pattern and sprouts with magnified view (inset, white dotted boundary) for hESC-ECs-seeded μV only constructs after 4 days of culture. Scale bar: 200 μm. (ii) μV + SA construct with perfused anastomotic connections. High magnification views of outlined regions 1 and 2 with corresponding in situ staining for mTm-hESC-ECs (DsRed+, red) and GFP-hESC-ECs (GFP+, green). Scale bars: 100 μm (left), 40 μm (right). Reprinted from Redd et al.57 BFP: blue fluorescent protein; EC: endothelial cell; ECM: extracellular matrix; EVT: extravillous trophoblast; GFP: green fluorescent protein; hESC-EC: human embryonic stem cell-derived endothelial cell; MSVT: multi-scale vascular tissue; SA: self-assembled.
| Study | Method | Cells used | Minimum diameter | Vascular network shape |
|---|---|---|---|---|
| Qiu et al. | Top-down EC lining method | HUVECs, HDMVECs, HLMVECs | ~20 μm | Cylindrical |
| de Graaf et al. | Top-down EC lining method | iPSC-ECs, HBVPs | ~300 μm | Cylindrical |
| Mandrycky et al. | Top-down EC lining method | HUVECs | ~200 μm | Spiral |
| Miller et al. | Top-down EC lining method | HUVECs, 10T1/2 cells | ~300 μm | Cylindrical |
| Yue et al. | Bottom-up self-assembly vascularisation method | ECFC-ECs, NHLFs | ~10 μm | Capillary |
| Nashimoto et al. | Bottom-up self-assembly vascularisation method | HUVECs, NHLFs | ~10 μm | Capillary |
| Phan et al. | Bottom-up self-assembly vascularisation method | ECFC-ECs, NHLFs | ~20 μm | Capillary |
| Tan et al. | Bottom-up self-assembly vascularisation method | HUVECs, NHLFs | ~20 μm | Capillary |
| Wang et al. | Hybrid method | ECFC-ECs, NHLFs | ~15 μm | Multi-scale vascular network |
| Park et al. | Hybrid method | HEMVECs, HLMVECs, HBMVECs, NHLFs | ~10 μm | Multi-scale vascular network |
| Debbi et al. | Hybrid method | HAMEC, DPSCs | ~20 μm | Multi-scale vascular network |
| Rebbi et al. | Hybrid method | hESC-ECs, HUVECs | ~20 μm | Multi-scale vascular network |
Table 1. Comparison of different in vitro vascularisation model construction methods
| Study | Method | Cells used | Minimum diameter | Vascular network shape |
|---|---|---|---|---|
| Qiu et al. | Top-down EC lining method | HUVECs, HDMVECs, HLMVECs | ~20 μm | Cylindrical |
| de Graaf et al. | Top-down EC lining method | iPSC-ECs, HBVPs | ~300 μm | Cylindrical |
| Mandrycky et al. | Top-down EC lining method | HUVECs | ~200 μm | Spiral |
| Miller et al. | Top-down EC lining method | HUVECs, 10T1/2 cells | ~300 μm | Cylindrical |
| Yue et al. | Bottom-up self-assembly vascularisation method | ECFC-ECs, NHLFs | ~10 μm | Capillary |
| Nashimoto et al. | Bottom-up self-assembly vascularisation method | HUVECs, NHLFs | ~10 μm | Capillary |
| Phan et al. | Bottom-up self-assembly vascularisation method | ECFC-ECs, NHLFs | ~20 μm | Capillary |
| Tan et al. | Bottom-up self-assembly vascularisation method | HUVECs, NHLFs | ~20 μm | Capillary |
| Wang et al. | Hybrid method | ECFC-ECs, NHLFs | ~15 μm | Multi-scale vascular network |
| Park et al. | Hybrid method | HEMVECs, HLMVECs, HBMVECs, NHLFs | ~10 μm | Multi-scale vascular network |
| Debbi et al. | Hybrid method | HAMEC, DPSCs | ~20 μm | Multi-scale vascular network |
| Rebbi et al. | Hybrid method | hESC-ECs, HUVECs | ~20 μm | Multi-scale vascular network |
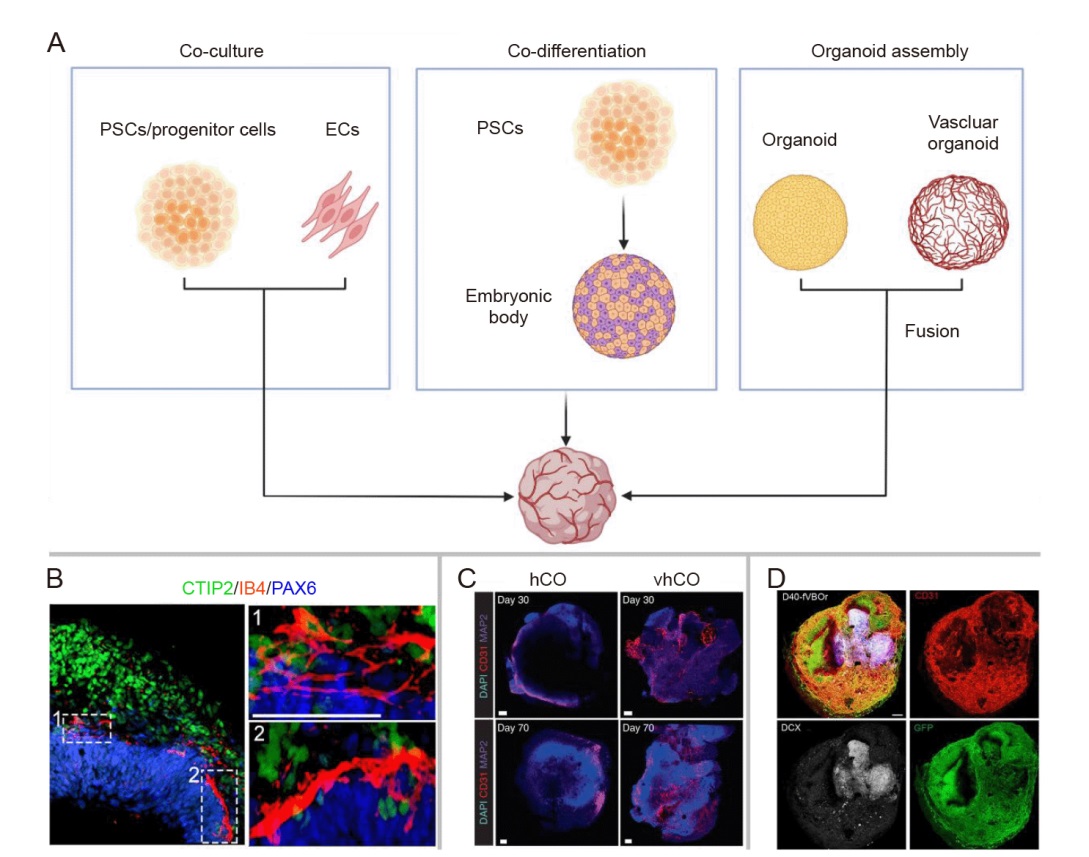
Figure 4. Strategies for organoid vascularisation. (A) Schematic representation of the principles underlying three strategies. Created with BioRender.com. (B) Representative immunofluorescence staining figure for CTIP2/IB4/PAX6 at day 65 to demonstrate that the vascular (IB4) structures would progressively extend into newborn neurons (CTIP2) with the development of vOrganoids. Scale bar: 50 μm. Reprinted from Shi et al.59 (C) Immunostaining of whole-mount vhCOs and control hCOs at different time points (days 30 and 70) for DAPI/CD31/MAP2. It illustrates that vhCOs had significantly more vessel area and length than control hCOs. Scale bars: 100 μm. Reprinted from Cakir et al.62 (D) Whole-mount staining results in the fused vasculature and brain organoids. D40-fVBOr represents the fused vasculature and brain organoids at day 40. Reprinted from Sun et al.65 Scale bar: 200 μm. CTIP2: COUP TF1 interacting protein 2; DAPI: 4′,6-diamidino-2-phenylindole; DCX: doublecortin; EC: endothelial cell; GFP: green fluorescent protein; hCOs: human cortical organoids; IB4: isolectin B4; MAP2: microtubule-associated protein 2; PAX6: paired box 6; PSC: pluripotent stem cell; vhCOs: vascularised human cortical organoids; vOrganoids: vascularised organoids.
| 1. | Riedl, A.; Schlederer, M.; Pudelko, K.; Stadler, M.; Walter, S.; Unterleuthner, D.; Unger, C.; Kramer, N.; Hengstschläger, M.; Kenner, L.; Pfeiffer, D.; Krupitza, G.; Dolznig, H. Comparison of cancer cells in 2D vs 3D culture reveals differences in AKT-mTOR-S6K signaling and drug responses. J Cell Sci. 2017, 130, 203-218. |
| 2. | Ekert, J. E.; Johnson, K.; Strake, B.; Pardinas, J.; Jarantow, S.; Perkinson, R.; Colter, D. C. Three-dimensional lung tumor microenvironment modulates therapeutic compound responsiveness in vitro--implication for drug development. PLoS One. 2014, 9, e92248. |
| 3. |
Li, Z.; Li, Q.; Zhou, C.; Lu, K.; Liu, Y.; Xuan, L.; Wang, X. Organoid-on-a-chip: current challenges, trends, and future scope toward medicine. Biomicrofluidics. 2023, 17, 051505.
doi: 10.1063/5.0171350 URL |
| 4. |
Quan, Y.; Sun, M.; Tan, Z.; Eijkel, J. C. T.; van den Berg, A.; van der Meer, A.; Xie, Y. Organ-on-a-chip: the next generation platform for risk assessment of radiobiology. RSC Adv. 2020, 10, 39521-39530.
doi: 10.1039/D0RA05173J URL |
| 5. |
Brancato, V.; Oliveira, J. M.; Correlo, V. M.; Reis, R. L.; Kundu, S. C. Could 3D models of cancer enhance drug screening? Biomaterials. 2020, 232, 119744.
doi: 10.1016/j.biomaterials.2019.119744 URL |
| 6. |
Takebe, T.; Zhang, B.; Radisic, M. Synergistic engineering: organoids meet organs-on-a-chip. Cell Stem Cell. 2017, 21, 297-300.
doi: 10.1016/j.stem.2017.08.016 URL |
| 7. | Zhu, Y.; Sun, L.; Wang, Y.; Cai, L.; Zhang, Z.; Shang, Y.; Zhao, Y. A biomimetic human lung-on-a-chip with colorful display of microphysiological breath. Adv Mater. 2022, 34, e2108972. |
| 8. |
Wang, L.; Tao, T.; Su, W.; Yu, H.; Yu, Y.; Qin, J. A disease model of diabetic nephropathy in a glomerulus-on-a-chip microdevice. Lab Chip. 2017, 17, 1749-1760.
doi: 10.1039/C7LC00134G URL |
| 9. |
Marsano, A.; Conficconi, C.; Lemme, M.; Occhetta, P.; Gaudiello, E.; Votta, E.; Cerino, G.; Redaelli, A.; Rasponi, M. Beating heart on a chip: a novel microfluidic platform to generate functional 3D cardiac microtissues. Lab Chip. 2016, 16, 599-610.
doi: 10.1039/C5LC01356A URL |
| 10. |
Crespo, M.; Vilar, E.; Tsai, S. Y.; Chang, K.; Amin, S.; Srinivasan, T.; Zhang, T.; Pipalia, N. H.; Chen, H. J.; Witherspoon, M.; Gordillo, M.; Xiang, J. Z.; Maxfield, F. R.; Lipkin, S.; Evans, T.; Chen, S. Colonic organoids derived from human induced pluripotent stem cells for modeling colorectal cancer and drug testing. Nat Med. 2017, 23, 878-884.
doi: 10.1038/nm.4355 URL |
| 11. |
Hassan, S.; Sebastian, S.; Maharjan, S.; Lesha, A.; Carpenter, A. M.; Liu, X.; Xie, X.; Livermore, C.; Zhang, Y. S.; Zarrinpar, A. Liver-on-a-chip models of fatty liver disease. Hepatology. 2020, 71, 733-740.
doi: 10.1002/hep.31106 URL |
| 12. |
Wu, Y.; Zhou, Y.; Paul, R.; Qin, X.; Islam, K.; Liu, Y. Adaptable microfluidic vessel-on-a-chip platform for investigating tumor metastatic transport in bloodstream. Anal Chem. 2022, 94, 12159-12166.
doi: 10.1021/acs.analchem.2c02556 URL |
| 13. |
Kilic, O.; Pamies, D.; Lavell, E.; Schiapparelli, P.; Feng, Y.; Hartung, T.; Bal-Price, A.; Hogberg, H. T.; Quinones-Hinojosa, A.; Guerrero-Cazares, H.; Levchenko, A. Brain-on-a-chip model enables analysis of human neuronal differentiation and chemotaxis. Lab Chip. 2016, 16, 4152-4162.
doi: 10.1039/C6LC00946H URL |
| 14. |
Lancaster, M. A.; Knoblich, J. A. Organogenesis in a dish: modeling development and disease using organoid technologies. Science. 2014, 345, 1247125.
doi: 10.1126/science.1247125 URL |
| 15. |
Wang, X.; Yamamoto, Y.; Wilson, L. H.; Zhang, T.; Howitt, B. E.; Farrow, M. A.; Kern, F.; Ning, G.; Hong, Y.; Khor, C. C.; Chevalier, B.; Bertrand, D.; Wu, L.; Nagarajan, N.; Sylvester, F. A.; Hyams, J. S.; Devers, T.; Bronson, R.; Lacy, D. B.; Ho, K. Y.; Crum, C. P.; McKeon, F.; Xian, W. Cloning and variation of ground state intestinal stem cells. Nature. 2015, 522, 173-178.
doi: 10.1038/nature14484 |
| 16. |
Nie, Y. Z.; Zheng, Y. W.; Miyakawa, K.; Murata, S.; Zhang, R. R.; Sekine, K.; Ueno, Y.; Takebe, T.; Wakita, T.; Ryo, A.; Taniguchi, H. Recapitulation of hepatitis B virus-host interactions in liver organoids from human induced pluripotent stem cells. EBioMedicine. 2018, 35, 114-123.
doi: 10.1016/j.ebiom.2018.08.014 URL |
| 17. | Li, L.; Knutsdottir, H.; Hui, K.; Weiss, M. J.; He, J.; Philosophe, B.; Cameron, A. M.; Wolfgang, C. L.; Pawlik, T. M.; Ghiaur, G.; Ewald, A. J.; Mezey, E.; Bader, J. S.; Selaru, F. M. Human primary liver cancer organoids reveal intratumor and interpatient drug response heterogeneity. JCI Insight. 2019, 4, e121490. |
| 18. |
Kretzschmar, K. Cancer research using organoid technology. J Mol Med (Berl). 2021, 99, 501-515.
doi: 10.1007/s00109-020-01990-z |
| 19. |
LeSavage, B. L.; Suhar, R. A.; Broguiere, N.; Lutolf, M. P.; Heilshorn, S. C. Next-generation cancer organoids. Nat Mater. 2022, 21, 143-159.
doi: 10.1038/s41563-021-01057-5 |
| 20. |
Yang, J.; Hirai, Y.; Iida, K.; Ito, S.; Trumm, M.; Terada, S.; Sakai, R.; Tsuchiya, T.; Tabata, O.; Kamei, K. I. Integrated-gut-liver-on-a-chip platform as an in vitro human model of non-alcoholic fatty liver disease. Commun Biol. 2023, 6, 310.
doi: 10.1038/s42003-023-04710-8 |
| 21. |
Ronaldson-Bouchard, K.; Teles, D.; Yeager, K.; Tavakol, D..; Zhao, Y.; Chramiec, A.; Tagore, S.; Summers, M.; Stylianos, S.; Tamargo, M.; Lee, B. M.; Halligan, S. P.; Abaci, E. H.; Guo, Z.; Jacków, J.; Pappalardo, A.; Shih, J.; Soni, R. K.; Sonar, S.; German, C.; Christiano, A. M.; Califano, A.; Hirschi, K. K.; Chen, C. S.; Przekwas, A.; Vunjak-Novakovic, G. NA multi-organ chip with matured tissue niches linked by vascular flow. Nat Biomed Eng. 2022, 6, 351-371.
doi: 10.1038/s41551-022-00882-6 |
| 22. |
Lee, H. R.; Sung, J. H. Multiorgan-on-a-chip for realization of gut-skin axis. Biotechnol Bioeng. 2022, 119, 2590-2601.
doi: 10.1002/bit.v119.9 URL |
| 23. |
Zhang, S.; Wan, Z.; Kamm, R. D. Vascularized organoids on a chip: strategies for engineering organoids with functional vasculature. Lab Chip. 2021, 21, 473-488.
doi: 10.1039/D0LC01186J URL |
| 24. |
Mandrycky, C. J.; Howard, C. C.; Rayner, S. G.; Shin, Y. J.; Zheng, Y. Organ-on-a-chip systems for vascular biology. J Mol Cell Cardiol. 2021, 159, 1-13.
doi: 10.1016/j.yjmcc.2021.06.002 URL |
| 25. |
Wang, X.; Sun, Q.; Pei, J. Microfluidic-based 3D engineered microvascular networks and their applications in vascularized microtumor models. Micromachines. 2018, 9, 493.
doi: 10.3390/mi9100493 URL |
| 26. |
Zhou, J.; Li, Y. S.; Chien, S. Shear stress-initiated signaling and its regulation of endothelial function. Arterioscler Thromb Vasc Biol. 2014, 34, 2191-2198.
doi: 10.1161/ATVBAHA.114.303422 URL |
| 27. |
Chistiakov, D. A.; Orekhov, A. N.; Bobryshev, Y. V. Effects of shear stress on endothelial cells: go with the flow. Acta Physiol (Oxf). 2017, 219, 382-408.
doi: 10.1111/apha.2017.219.issue-2 URL |
| 28. | Baeyens, N.; Bandyopadhyay, C.; Coon, B. G.; Yun, S.; Schwartz, M. A. Endothelial fluid shear stress sensing in vascular health and disease. J Clin Invest. 2016, 126, 821-828. |
| 29. |
Wu, J.; Shadden, S. C. Coupled simulation of hemodynamics and vascular growth and remodeling in a subject-specific geometry. Ann Biomed Eng. 2015, 43, 1543-1554.
doi: 10.1007/s10439-015-1287-6 URL |
| 30. |
Homan, K. A.; Gupta, N.; Kroll, K. T.; Kolesky, D. B.; Skylar-Scott, M.; Miyoshi, T.; Mau, D.; Valerius, M. T.; Ferrante, T.; Bonventre, J. V.; Lewis, J. A.; Morizane, R. Flow-enhanced vascularization and maturation of kidney organoids in vitro. Nat Methods. 2019, 16, 255-262.
doi: 10.1038/s41592-019-0325-y |
| 31. |
Kramer, R. H.; Nicolson, G. L. Interactions of tumor cells with vascular endothelial cell monolayers: a model for metastatic invasion. Proc Natl Acad Sci U S A. 1979, 76, 5704-5708.
doi: 10.1073/pnas.76.11.5704 URL |
| 32. |
Young, E. W.; Watson, M. W.; Srigunapalan, S.; Wheeler, A. R.; Simmons, C. A. Technique for real-time measurements of endothelial permeability in a microfluidic membrane chip using laser-induced fluorescence detection. Anal Chem. 2010, 82, 808-816.
doi: 10.1021/ac901560w URL |
| 33. |
Halldorsson, S.; Lucumi, E.; Gómez-Sjöberg, R.; Fleming, R. M. T. Advantages and challenges of microfluidic cell culture in polydimethylsiloxane devices. Biosens Bioelectron. 2015, 63, 218-231.
doi: 10.1016/j.bios.2014.07.029 URL |
| 34. |
Hasan, A.; Paul, A.; Vrana, N. E.; Zhao, X.; Memic, A.; Hwang, Y. S.; Dokmeci, M. R.; Khademhosseini, A. Microfluidic techniques for development of 3D vascularized tissue. Biomaterials. 2014, 35, 7308-7325.
doi: 10.1016/j.biomaterials.2014.04.091 URL |
| 35. | Li, X.; Xu, S.; He, P.; Liu, Y. In vitro recapitulation of functional microvessels for the study of endothelial shear response, nitric oxide and [Ca2+]i. PLoS One. 2015, 10, e0126797. |
| 36. | Huynh, J.; Nishimura, N.; Rana, K.; Peloquin, J. M.; Califano, J. P.; Montague, C. R.; King, M. R.; Schaffer, C. B.; Reinhart-King, C. A. Age-related intimal stiffening enhances endothelial permeability and leukocyte transmigration. Sci Transl Med. 2011, 3, 112ra122. |
| 37. |
Qiu, Y.; Ahn, B.; Sakurai, Y.; Hansen, C. E.; Tran, R.; Mimche, P. N.; Mannino, R. G.; Ciciliano, J. C.; Lamb, T. J.; Joiner, C. H.; Ofori-Acquah, S. F.; Lam, W. A. Microvasculature-on-a-chip for the long-term study of endothelial barrier dysfunction and microvascular obstruction in disease. Nat Biomed Eng. 2018, 2, 453-463.
doi: 10.1038/s41551-018-0224-z |
| 38. |
de Graaf, M. N. S.; Cochrane, A.; van den Hil, F. E.; Buijsman, W.; van der Meer, A. D.; van den Berg, A.; Mummery, C. L.; Orlova, V. V. Scalable microphysiological system to model three-dimensional blood vessels. APL Bioeng. 2019, 3, 026105.
doi: 10.1063/1.5090986 URL |
| 39. | Wong, K. H.; Truslow, J. G.; Khankhel, A. H.; Chan, K. L.; Tien, J. Artificial lymphatic drainage systems for vascularized microfluidic scaffolds. J Biomed Mater Res A. 2013, 101, 2181-2190. |
| 40. | Mandrycky, C.; Hadland, B.; Zheng, Y. 3D curvature-instructed endothelial flow response and tissue vascularization. Sci Adv. 2020, 6, eabb3629. |
| 41. | Saggiomo, V.; Velders, A H. Simple 3D printed scaffold-removal method for the fabrication of intricate microfluidic devices. Adv Sci (Weinh). 2015, 2, 1500125. |
| 42. |
Miller, J. S.; Stevens, K. R.; Yang, M. T.; Baker, B. M.; Nguyen, D. H.; Cohen, D. M.; Toro, E.; Chen, A. A.; Galie, P. A.; Yu, X.; Chaturvedi, R.; Bhatia, S. N.; Chen, C. S. Rapid casting of patterned vascular networks for perfusable engineered three-dimensional tissues. Nat Mater. 2012, 11, 768-774.
doi: 10.1038/nmat3357 |
| 43. |
Shirure, V. S.; Hughes, C. C. W.; George, S. C. Engineering vascularized organoid-on-a-chip models. Annu Rev Biomed Eng. 2021, 23, 141-167.
doi: 10.1146/bioeng.2021.23.issue-1 URL |
| 44. |
Whisler, J. A.; Chen, M. B.; Kamm, R. D. Control of perfusable microvascular network morphology using a multiculture microfluidic system. Tissue Eng Part C Methods. 2014, 20, 543-552.
doi: 10.1089/ten.tec.2013.0370 URL |
| 45. | Takakura, N. Role of intimate interactions between endothelial cells and the surrounding accessory cells in the maturation of blood vessels. J Thromb Haemost. 2011, 9 Suppl 1, 144-150. |
| 46. |
Shepherd, B. R.; Jay, S. M.; Saltzman, W. M.; Tellides, G.; Pober, J. S. Human aortic smooth muscle cells promote arteriole formation by coengrafted endothelial cells. Tissue Eng Part A. 2009, 15, 165-173.
doi: 10.1089/ten.tea.2008.0010 URL |
| 47. |
Koike, N.; Fukumura, D.; Gralla, O.; Au, P.; Schechner, J. S.; Jain, R. K. Tissue engineering: creation of long-lasting blood vessels. Nature. 2004, 428, 138-139.
doi: 10.1038/428138a |
| 48. |
Logsdon, E. A.; Finley, S. D.; Popel, A. S.; Mac Gabhann, F. A systems biology view of blood vessel growth and remodelling. J Cell Mol Med. 2014, 18, 1491-1508.
doi: 10.1111/jcmm.2014.18.issue-8 URL |
| 49. |
Zhao, Y.; Adjei, A. A. Targeting angiogenesis in cancer therapy: moving beyond vascular endothelial growth factor. Oncologist. 2015, 20, 660-673.
doi: 10.1634/theoncologist.2014-0465 URL |
| 50. |
Yue, T.; Zhao, D.; Phan, D. T. T.; Wang, X.; Park, J. J.; Biviji, Z.; Hughes, C. C. W.; Lee, A. P. A modular microfluidic system based on a multilayered configuration to generate large-scale perfusable microvascular networks. Microsyst Nanoeng. 2021, 7, 4.
doi: 10.1038/s41378-020-00229-8 |
| 51. |
Nashimoto, Y.; Hayashi, T.; Kunita, I.; Nakamasu, A.; Torisawa, Y. S.; Nakayama, M.; Takigawa-Imamura, H.; Kotera, H.; Nishiyama, K.; Miura, T.; Yokokawa, R. Integrating perfusable vascular networks with a three-dimensional tissue in a microfluidic device. Integr Biol (Camb). 2017, 9, 506-518.
doi: 10.1039/C7IB00024C URL |
| 52. |
Phan, D. T. T.; Wang, X.; Craver, B. M.; Sobrino, A.; Zhao, D.; Chen, J. C.; Lee, L. Y. N.; George, S. C.; Lee, A. P.; Hughes, C. C. W. A vascularized and perfused organ-on-a-chip platform for large-scale drug screening applications. Lab Chip. 2017, 17, 511-520.
doi: 10.1039/C6LC01422D URL |
| 53. |
Tan, S. Y.; Jing, Q.; Leung, Z.; Xu, Y.; Cheng, L. K. W.; Tam, S. S. T.; Wu, A. R. Transcriptomic analysis of 3D vasculature-on-a-chip reveals paracrine factors affecting vasculature growth and maturation. Lab Chip. 2022, 22, 3885-3897.
doi: 10.1039/D2LC00570K URL |
| 54. |
Wang, X.; Phan, D. T.; Sobrino, A.; George, S. C.; Hughes, C. C.; Lee, A. P. Engineering anastomosis between living capillary networks and endothelial cell-lined microfluidic channels. Lab Chip. 2016, 16, 282-290.
doi: 10.1039/C5LC01050K URL |
| 55. |
Park, J. Y.; Mani, S.; Clair, G.; Olson, H. M.; Paurus, V. L.; Ansong, C. K.; Blundell, C.; Young, R.; Kanter, J.; Gordon, S.; Yi, A. Y.; Mainigi, M.; Huh, D. D. A microphysiological model of human trophoblast invasion during implantation. Nat Commun. 2022, 13, 1252.
doi: 10.1038/s41467-022-28663-4 |
| 56. |
Debbi, L.; Zohar, B.; Shuhmaher, M.; Shandalov, Y.; Goldfracht, I.; Levenberg, S. Integrating engineered macro vessels with self-assembled capillaries in 3D implantable tissue for promoting vascular integration in-vivo. Biomaterials. 2022, 280, 121286.
doi: 10.1016/j.biomaterials.2021.121286 URL |
| 57. |
Redd, M. A.; Zeinstra, N.; Qin, W.; Wei, W.; Martinson, A.; Wang, Y.; Wang, R. K.; Murry, C. E.; Zheng, Y. Patterned human microvascular grafts enable rapid vascularization and increase perfusion in infarcted rat hearts. Nat Commun. 2019, 10, 584.
doi: 10.1038/s41467-019-08388-7 |
| 58. |
Takebe, T.; Sekine, K.; Enomura, M.; Koike, H.; Kimura, M.; Ogaeri, T.; Zhang, R. R.; Ueno, Y.; Zheng, Y. W.; Koike, N.; Aoyama, S.; Adachi, Y.; Taniguchi, H. Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature. 2013, 499, 481-484.
doi: 10.1038/nature12271 |
| 59. | Shi, Y.; Sun, L.; Wang, M.; Liu, J.; Zhong, S.; Li, R.; Li, P.; Guo, L.; Fang, A.; Chen, R.; Ge, W. P.; Wu, Q.; Wang, X. Vascularized human cortical organoids (vOrganoids) model cortical development in vivo. PLoS Biol. 2020, 18, e3000705. |
| 60. |
Salmon, I.; Grebenyuk, S.; Abdel Fattah, A. R.; Rustandi, G.; Pilkington, T.; Verfaillie, C.; Ranga, A. Engineering neurovascular organoids with 3D printed microfluidic chips. Lab Chip. 2022, 22, 1615-1629.
doi: 10.1039/D1LC00535A URL |
| 61. | Wan, Z.; Floryan, M. A.; Coughlin, M. F.; Zhang, S.; Zhong, A. X.; Shelton, S. E.; Wang, X.; Xu, C.; Barbie, D. A.; Kamm, R. D. New strategy for promoting vascularization in tumor spheroids in a microfluidic assay. Adv Healthc Mater. 2023, 12, e2201784. |
| 62. |
Cakir, B.; Xiang, Y.; Tanaka, Y.; Kural, M. H.; Parent, M.; Kang, Y. J.; Chapeton, K.; Patterson, B.; Yuan, Y.; He, C. S.; Raredon, M. S. B.; Dengelegi, J.; Kim, K. Y.; Sun, P.; Zhong, M.; Lee, S.; Patra, P.; Hyder, F.; Niklason, L. E.; Lee, S. H.; Yoon, Y. S.; Park, I. H. Engineering of human brain organoids with a functional vascular-like system. Nat Methods. 2019, 16, 1169-1175.
doi: 10.1038/s41592-019-0586-5 |
| 63. |
Takasato, M.; Er, P. X.; Chiu, H. S.; Maier, B.; Baillie, G. J.; Ferguson, C.; Parton, R. G.; Wolvetang, E. J.; Roost, M. S.; Chuva de Sousa Lopes, S. M.; Little, M. H. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature. 2015, 526, 564-568.
doi: 10.1038/nature15695 |
| 64. |
Song, L.; Yuan, X.; Jones, Z.; Griffin, K.; Zhou, Y.; Ma, T.; Li, Y. Assembly of human stem cell-derived cortical spheroids and vascular spheroids to model 3-D brain-like tissues. Sci Rep. 2019, 9, 5977.
doi: 10.1038/s41598-019-42439-9 |
| 65. | Sun, X. Y.; Ju, X. C.; Li, Y.; Zeng, P. M.; Wu, J.; Zhou, Y. Y.; Shen, L. B.; Dong, J.; Chen, Y. J.; Luo, Z. G. Generation of vascularized brain organoids to study neurovascular interactions. Elife. 2022, 11, e76707. |
| 66. |
Li, M.; Gao, L.; Zhao, L.; Zou, T.; Xu, H. Toward the next generation of vascularized human neural organoids. Med Res Rev. 2023, 43, 31-54.
doi: 10.1002/med.v43.1 URL |
| 67. |
Zhao, X.; Xu, Z.; Xiao, L.; Shi, T.; Xiao, H.; Wang, Y.; Li, Y.; Xue, F.; Zeng, W. Review on the vascularization of organoids and organoids-on-a-chip. Front Bioeng Biotechnol. 2021, 9, 637048.
doi: 10.3389/fbioe.2021.637048 URL |
| 68. |
LaMontagne, E.; Muotri, A. R.; Engler, A. J. Recent advancements and future requirements in vascularization of cortical organoids. Front Bioeng Biotechnol. 2022, 10, 1048731.
doi: 10.3389/fbioe.2022.1048731 URL |
| 69. |
Liu, H.; Zhang, X.; Liu, J.; Qin, J. Vascularization of engineered organoids. BMEMat. 2023, 1, e12031.
doi: 10.1002/bmm2.v1.3 URL |
| 70. |
Sato, Y.; Asahi, T.; Kataoka, K. Integrative single-cell RNA-seq analysis of vascularized cerebral organoids. BMC Biol. 2023, 21, 245.
doi: 10.1186/s12915-023-01711-1 |
| 71. |
Nashimoto, Y.; Okada, R.; Hanada, S.; Arima, Y.; Nishiyama, K.; Miura, T.; Yokokawa, R. Vascularized cancer on a chip: The effect of perfusion on growth and drug delivery of tumor spheroid. Biomaterials. 2020, 229, 119547.
doi: 10.1016/j.biomaterials.2019.119547 URL |
| 72. |
van den Berg, C. W.; Ritsma, L.; Avramut, M. C.; Wiersma, L. E.; van den Berg, B. M.; Leuning, D. G.; Lievers, E.; Koning, M.; Vanslambrouck, J. M.; Koster, A. J.; Howden, S. E.; Takasato, M.; Little, M. H.; Rabelink, T. J. Renal subcapsular transplantation of PSC-derived kidney organoids induces neo-vasculogenesis and significant glomerular and tubular maturation in vivo. Stem Cell Reports. 2018, 10, 751-765.
doi: 10.1016/j.stemcr.2018.01.041 URL |
| [1] | Chen-Hui Mi, Xin-Ya Qi, Yan-Wen Ding, Jing Zhou, Jin-Wei Dao, Dai-Xu Wei. Recent advances of medical polyhydroxyalkanoates in musculoskeletal system [J]. Biomaterials Translational, 2023, 4(4): 234-247. |
| [2] | Ross H. McWilliam, Wenlong Chang, Zhao Liu, Jiayuan Wang, Fengxuan Han, Richard A. Black, Junxi Wu, Xichun Luo, Bin Li, Wenmiao Shu. Three-dimensional biofabrication of nanosecond laser micromachined nanofibre meshes for tissue engineered scaffolds [J]. Biomaterials Translational, 2023, 4(2): 104-114. |
| [3] | Ahlam A. Abdalla, Catherine J. Pendegrass. Biological approaches to the repair and regeneration of the rotator cuff tendon-bone enthesis: a literature review [J]. Biomaterials Translational, 2023, 4(2): 85-103. |
| [4] | Rob Jess, Tao Ling, Yi Xiong, Chris J. Wright, Feihu Zhao. Mechanical environment for in vitro cartilage tissue engineering assisted by in silico models [J]. Biomaterials Translational, 2023, 4(1): 18-26. |
| [5] | Chavee Laomeephol, Helena Ferreira, Sorada Kanokpanont, Jittima Amie Luckanagul, Nuno M Neves, Siriporn Damrongsakkul. Osteogenic differentiation of encapsulated cells in dexamethasone–loaded phospholipid–induced silk fibroin hydrogels [J]. Biomaterials Translational, 2022, 3(3): 213-220. |
| [6] | Ricardo Donate, Maryam Tamaddon, Viviana Ribeiro, Mario Monzón, J. Miguel Oliveira, Chaozong Liu. Translation through collaboration: practice applied in BAMOS project in in vivo testing of innovative osteochondral scaffolds [J]. Biomaterials Translational, 2022, 3(2): 102-104. |
| [7] | Melika Sahranavard, Soulmaz Sarkari, SeyedehMina Safavi, Farnaz Ghorbani. Three-dimensional bio-printing of decellularized extracellular matrix-based bio-inks for cartilage regeneration: a systematic review [J]. Biomaterials Translational, 2022, 3(2): 105-115. |
| [8] | Emma Steijvers, Armaan Ghei, Zhidao Xia. Manufacturing artificial bone allografts: a perspective [J]. Biomaterials Translational, 2022, 3(1): 65-80. |
| [9] | Ke Hu, Yuxuan Li, Zunxiang Ke, Hongjun Yang, Chanjun Lu, Yiqing Li, Yi Guo, Weici Wang. History, progress and future challenges of artificial blood vessels: a narrative review [J]. Biomaterials Translational, 2022, 3(1): 81-98. |
| [10] | Yizhong Peng, Jinye Li, Hui Lin, Shuo Tian, Sheng Liu, Feifei Pu, Lei Zhao, Kaige Ma, Xiangcheng Qing, Zengwu Shao. Endogenous repair theory enriches construction strategies for orthopaedic biomaterials: a narrative review [J]. Biomaterials Translational, 2021, 2(4): 343-360. |
| [11] | Xirui Jing, Qiuyue Ding, Qinxue Wu, Weijie Su, Keda Yu, Yanlin Su, Bing Ye, Qing Gao, Tingfang Sun, Xiaodong Guo. Magnesium-based materials in orthopaedics: material properties and animal models [J]. Biomaterials Translational, 2021, 2(3): 197-213. |
| [12] | Yiqing Wang, Xiangyu Chu, Bing Wang. Recombinant adeno-associated virus-based gene therapy combined with tissue engineering for musculoskeletal regenerative medicine [J]. Biomaterials Translational, 2021, 2(1): 19-29. |
| [13] | Maryam Tamaddon, Helena Gilja, Ling Wang, J. Miguel Oliveira, Xiaodan Sun, Rongwei Tan, Chaozong Liu. Osteochondral scaffolds for early treatment of cartilage defects in osteoarthritic joints: from bench to clinic [J]. Biomaterials Translational, 2020, 1(1): 3-17. |
| [14] | Xing Yang, Yuanyuan Li, Xujie Liu, Wei He, Qianli Huang, Qingling Feng. Nanoparticles and their effects on differentiation of mesenchymal stem cells [J]. Biomaterials Translational, 2020, 1(1): 58-68. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||